What you can read in here is a set of my loose notes on complex systems and biology. I want to learn about the topic as fast as I can, so if I’m wrong anywhere, please point that to me. This post is an overview and indication of issues I’d like to cover.
Complex adaptive systems (CAS) are the heart of many phenomenas we observe every day, such as global trade, ecosystems, human body, immune system, internet and even language. Complexity of CAS does not equall to amount of information, rather it’s a indication of complex, positive and negative interactions of its components. All CAS feature a common set of dualisms:
- distinct/connected – CAS are built of a large number of agents that interact simultaneously and independently but all together become tightly regulated system (other names: individual/system or distributed/collective)
- robust/sensitive – CAS are pretty robust, yet at the same time are quite sensitive to initial conditions and some signals (see butterfly effect); both features are unpredictable
- local/global – protein is a CAS, protein network is a CAS, cell is a CAS, tissue is a CAS, organism is a CAS, society is a CAS; agents of a CAS, can be CAS themselves
- adaptive/evolving – CAS is able to adapt as a system and usually its agents are also mutually adaptive, and at the same time CAS is evolving; even if local landscape prefers simpler solutions (adaptation) CAS usually evolve toward bigger complexity
These dualisms are in some sense as artificial as wave-particle dualism. Complex system has all these features at the same time – their visibility depends only on design of a experiment. As a result, CAS present a common set of features: they are self-organizing, coherent, emergent and non-linear.
Probably the best so far representation of CAS is a network, which has a number of important features: it is scale-free (distribution of links in the network tends to follow power law), clustered (“friend of my friend is likely my friend too”) and small-world-like (diameter of a network is small, aka “six degrees of separation”). Such representation has been applied to biological complex systems, such as metabolic networks, or protein-protein interaction networks with a great success. However please remember that it’s only representation and many times people argued that scale-free networks may not be the best approximation of natural networks (see for example this recent paper).
Scale-free or not, network representation doesn’t address all dualities mentioned above, especially last two. Naturally emerging levels of organisation and relation between adaptation and evolution of complex systems are rarely studied from biological point of view, probably because we don’t have a clear idea how to reduce these phenomenas to something measurable.
In the next posts, I will try to cover other CAS representations and computational approaches to CAS modeling.


![Reblog this post [with Zemanta]](https://i0.wp.com/img.zemanta.com/reblog_e.png)


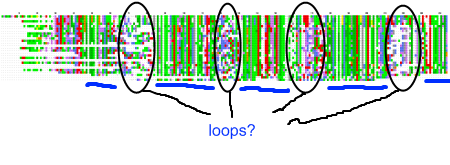 My recent post on visual analytics in bioinformatics lacked a specific example, but I’m happy to finally provide one (happiness comes also from the fact that respective publication is finally in press). The image above shows a multiple pairwise alignment from BLAST of a putative inner membrane protein from
My recent post on visual analytics in bioinformatics lacked a specific example, but I’m happy to finally provide one (happiness comes also from the fact that respective publication is finally in press). The image above shows a multiple pairwise alignment from BLAST of a putative inner membrane protein from 






