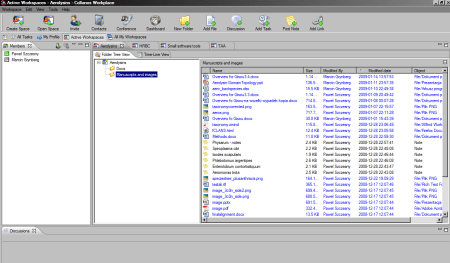
One of my workspaces in Collanos
For some time already I was looking for a tool that would eliminate a need for sending files back and forth between people collaborating on a the same project. While I’m perfectly aware of various solutions such as wikis, version control systems or online office suites, I didn’t feel like I could convince my collaborators to use any of these. One of the reasons is always a feeling of insecurity when using publicly hosted platform (BTW, this is not that uncommon among scientists – I know at least one scientific institution in Western Europe that explicitly forbids using Google apps, especially Gmail for work-related stuff, because of Google’s privacy policy). The other reason was that such solutions are not the best choice when working on binary files (most of my projects do not involve collaborative programming). When I stumbled across Collanos Workplace, which offers peer-to-peer synchronization (although without revision control), instead of a central-server based, I’ve decided to give it a try. For the last couple of weeks I’ve been using Collanos to collaborate on one relatively simple project and the experience was quite positive.
At first, I thought that Collanos may serve mainly as a tool for secure peer-to-peer files sharing with an information who changed what etc. It turned out that this is a capable project management application, that has a chat and discussion panel, one can post notes, links add tasks and assign them to team members. Files are stored is a separate directory – after one adds a file to Collanos, it should be opened from the application, not from original folder. This seemed a mistake in design at first, but I appreciated it very quickly. Synchronization of project directory would mean sharing all of its contents and that can be sometimes in the range of many GBs. From time to time some bug appeared here or there, but overall it worked as expected. Peer-to-peer sharing means that both people have to be online for synchronization, but so far situation that I switched computer off before a person could download my changes happened only once and it was during a weekend.
As a side note, it’s nice to see that Eclipse becomes an application platform for quite a number of programs. See for example this list of Eclipse-based software.
![Reblog this post [with Zemanta]](https://i0.wp.com/img.zemanta.com/reblog_e.png)






Manual sequence analysis – some common mistakes
This is a topic I probably will come back to on many occasions. Publication with very wrong sequence analysis like the one Stephen Spiro pointed out on his blog is not an exception. I may agree that large scale analysis can stand quick and dirty treatment of protein sequence (and some error propagation at the same time). In large scale analysis nobody cares if the domain assignment is 100% right (it isn’t), if there are false positives (there are) or even if the material to begin with (protein sequences for example) is free of errors (it is not) – as long as the overall quality of the work is acceptable. However, this optimistic approach cannot be applied to the manual protein sequence analysis. Simply errors introduced in such cases are a way more important. How to avoid some of these errors? A few common mistakes that come to my mind are:
It’s so far all I could think of. Do you have any other suggestions? Let me know.
Posted by Pawel Szczesny on September 25, 2007 in Comments, Research, Research skills