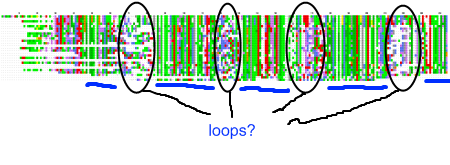
 My recent post on visual analytics in bioinformatics lacked a specific example, but I’m happy to finally provide one (happiness comes also from the fact that respective publication is finally in press). The image above shows a multiple pairwise alignment from BLAST of a putative inner membrane protein from Porphyromonas gingivalis. Image is small but it does not really matter – colour patches seem to be visible anyway.
My recent post on visual analytics in bioinformatics lacked a specific example, but I’m happy to finally provide one (happiness comes also from the fact that respective publication is finally in press). The image above shows a multiple pairwise alignment from BLAST of a putative inner membrane protein from Porphyromonas gingivalis. Image is small but it does not really matter – colour patches seem to be visible anyway.
Regions marked with ovals are clearly less conserved, than other part of the protein. There are five hydrophobic (green patches, underlined with blue lines) regions in this alignment (I ignore N-terminus, as this is likely the signal peptide), however the three inner ones appear to be of similar length, while the outer ones seem to be of the half as long as the inner ones. If we assume that the single unit is the short one, we can summarize the protein as follows: 8 beta structures, four long loops, for short loops. It looks like an eight-stranded outer membrane beta-barrel. Almost structure prediction, but without a structure.
I could end the story here, but the model didn’t fit previously published data. Its localization in the inner membrane was confirmed by an experiment, however pores in the inner membrane are considered very harmfull 😉 . Fortunately, one of my colleagues explained to me that particular localization technique is not 100% reliable, so I gathered more evidence, created detailed description of topology and the other group has designed experiments which confirmed my visual analysis.
Lessons learned? Maybe without this feedback on quality of that experimental technique, I would still claim that this is OM beta-barrel. Or maybe not. But I’ve learned that to safely ignore experimental results, one needs a more than a intuition. Also, it shows that sometimes looking at the results, is all one needs to make a reasonable prediction (I still have no idea what were E-values of these BLAST hits, but does it matter?).
![Reblog this post [with Zemanta]](https://i0.wp.com/img.zemanta.com/reblog_e.png)






Brad Chapman
February 5, 2009 at 01:15
Pawel;
This is great and inspired a bit of coding on my part to calculate and display conservation across a protein based on BLAST results. When I am putting together a post with the code, it would be great to have a graph of your protein displayed here. Is the protein ID hush-hush, or can you share it? If not yet, I can use a favorite gene. Thanks,
Brad
Marcin Cieslik
February 5, 2009 at 01:36
Brad you could reverse-engineer the PSSM from this low-res image. No Ids are hush-hush enough;)
Pawel Szczesny
February 5, 2009 at 10:01
Brad, of course I can share it: it’s this protein.
Marcin, assuming that you make no assumption on color codes (if you can guess the code, it’s trivial), how would you proceed?
Marcin Cieslik
February 6, 2009 at 03:59
(assuming I have been given only the image)
My first thought was to:
– screen alignment vis software and find this color code
– decode colors to residue groups
– based on the length of the non-loop fragments and their amphipaticity decide if they are more likely alpha-helices or beta-strands
– I would reconstruct each site in the PSSM by sampling the residues from the groups which make up the columns according to the number of residues in the MSA which are of this group and some general alpha-helix or beta-strand propensities
– if this would not work I would further try to refine the PSSMs for different organisms (e.g. bacteria, archea etc. and using their amino acid frequencies) and/or specific protein types.
If I didn’t know the color-coding… hmmm… The number of possible 7?8?-color reduced alphabets from the 20 amino acids is enormous, but on the other hand most of them should be so lousy that no psi-blast queries would pop-up. My idea would be to break this task down to a) breaking the alphabet b) finding the protein with probably more then one reduced alphabet being blasted. Still this is an alignment so it has to be somehow optimal given some substitution matrix and insertion/deletion model. I would try to find the alphabet which maximises the likelihood of producing the alignment given commonly used substitution matrices. Honestly I don’t know how to sample/converge alphabets. Probably I would start by trying to _really_ understand how joint alignment/phylogeny given evolution model: http://www.biomath.ucla.edu/msuchard/bali-phy/Redelings_and_Suchard_2005.pdf and how to estimate Q-matrices from alignment/phylogeny using MCMC. (lost the reference for this one will fix it)
Andrew Perry
February 7, 2009 at 01:50
Reading the first part of this post, I thought you were about to announce a ‘paradigm busting’ finding of an integral inner membrane beta-barrel protein ! Then again, after the structure of Wza was revealed (apparently a helical integral outer membrane protein), I guess a beta-barrel in the inner membrane couldn’t be entirely ruled out. For instance … some of the smaller 8-stranded barrels often don’t show much ion passage – maybe a non-pore forming one could exist in the inner membrane without causing too much havoc ?
Despite that, there is no known protein import machinery specialized for beta-barrel insertion into the inner membrane, so it seems unlikely (based on what we know, the transmembrane strands wouldn’t be hydrophobic enough for SecYEG dependent integration into the membrane, and spontaneous insertion can happen in vitro but is too slow for bacteria).
Anyhow, nice post – it’s a good example of how useful the interplay of sequence analysis and wet-lab experiments can work as ‘sanity checks’ on experimental results that may otherwise be taken as gospel.
Brad Chapman
February 8, 2009 at 00:32
Pawel;
Awesome. Thanks for the pointer; it was great to be able to sanity check against your by eye findings. I put together an automated script inspired by this with an alternative visualization of conserved regions. The post on the code is here.
Marcin — wow, and here I thought you were joking. It turns out you are a master at deconstructing alignment figures. I will be certain to keep all my top secret pictures extra blurry from now on.
Ana Rojas
February 8, 2009 at 12:39
Hi Pawel!
Nice post. Definitely to get insights into real biology experimental sicence data shouldn´t be ruled out, but for sure they shouldn´t be considered as standard of truth.
Keep on going.