Almost a year ago I wrote a post about officially becoming “freelance scientist”. I didn’t really know what I was doing, but taking over the world from a small flat in the middle of nowhere in Poland sounded like a good idea. And it definitely was a good idea, however not in a way I thought it would be. Today I am hoping that all things will go fine and I’ll be employed since February in a calm academic environment.
What I aimed for?
My plan was to become freelance scientist – to be able to thrive financially and intelectually without relying on gaming grant systems. I had hoped to secure support from private sources, form a virtual institute and live happily in the middle of nowhere while still having an impact on the world’s science, possibly all under “open research” badge.
What didn’t work?
Side comment: if you read excellent piece by Hugh MacLeod entitled “How to be creative”, you shouldn’t find the issues below surprising :).
The main reason I started to look for a job already some weeks ago was that freelancing as a scientist turned out to be unsustainable financially. And don’t get me wrong – money wasn’t an issue, as long as I was willing to put all my time into other’s people projects. All. My. Time. Booking some time to work on my own ideas meant burning savings at a quick rate. But I had to work on my own ideas. I didn’t feel like I’m learning very much, because I worked on things I was already quite good at. Intellectual stretching was not that big.
Because of the issue above, I’ve put together far less work than I aimed to. I have lots of posts, manuscripts and presentations which I didn’t have time to finish. I was too busy doing freelance work, finishing the projects I had promised to do, inventing new projects and hiding under a bed worrying about where this is all going.
Working in the middle of nowhere was a plain mistake. It sounds nice, but f2f networking (“showing up”) is far more important than I’ve thought. Working in Poland is an issue on its own (no matter if you’re a freelancing or an academic scientist); working outside of any major city makes it even worse.
Partially connected to the former issue was the fact that I tried to do all things alone. Wrong. Very wrong. Things like virtual institute will not work, unless there’s a team. Period.
And finally, I didn’t give myself enough time to make the whole system work. It turned out that I had no idea about so many things influencing money-flow in the system, that it’s not surprising at all that it didn’t click in so short (12 months) time.
What worked?
One of two biggest advantages of this crazy 12 months was that it was a great learning experience. When I look at my older colleagues working in academic environment, I’m pretty sure they don’t experience “felling like an idiot” moments all that often. Such moments happen quite frequently in grad school, but seem to become rarer the further science career advances. On a contrary, I had such moments all the time in the last year. I was experimenting with blog posts, stupid ideas, unbalanced opinions and I was scared as hell each time. And I have learnt much more than I would do playing safe. Have you watched Ken Robinson’s talk at TED? He put a beatiful phrase – “prepared to be wrong”. Keep that in mind.
People were second most important factor here. I was amazed by a number of people that have helped me along the way. Lots of them have encouraged me, pointed to useful resources or invested significant amount of time into answering my silly questions. Many times I was blown away by the help I had not expected. Biogang/Life Scientists community rules.
Frequently quoted phrase from Bill Hooker’s essay, “I’ve never had an idea that couldn’t be improved by sharing it with as many people as possible — and I don’t think anyone else has, either.”, turned out to be absolutely true. Each time I presented my ideas, people were interacting with them, not judging them. I was given suggestions I would not come up with by myself, even if it was clear that we’re not going to do business together.
What now?
The job I hope to land next year is going to address the things I’ve written about above. I hope to have some financial stability and necessary time to advance my plans. It will also provide a support for such events like “Startup weekend in science”, which I plan to invite you all later next year.
The main goal is still valid and I don’t give up on it. I’ve found (or rather the other way round) a real-life example, ProTech Institute from Lithuania, which means that it can be done – it’s just a little harder if you’re a (still, but not for long) PhD student.
So it’s end of freelancing for now. Lessons learned. Back to real-life™ again :).
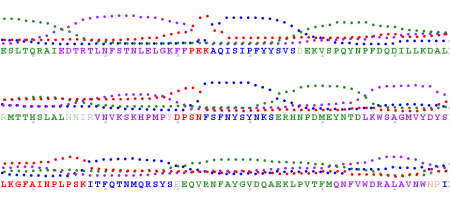
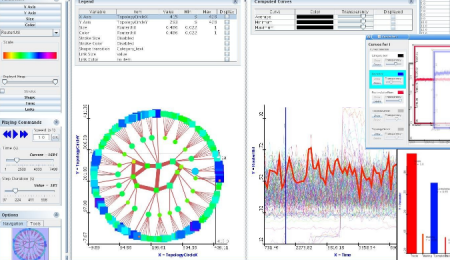
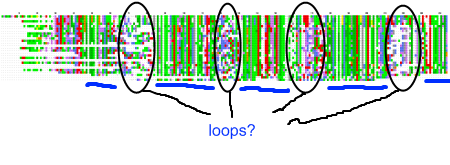 My recent post on visual analytics in bioinformatics lacked a specific example, but I’m happy to finally provide one (happiness comes also from the fact that respective publication is finally in press). The image above shows a multiple pairwise alignment from BLAST of a putative inner membrane protein from Porphyromonas gingivalis. Image is small but it does not really matter – colour patches seem to be visible anyway.
My recent post on visual analytics in bioinformatics lacked a specific example, but I’m happy to finally provide one (happiness comes also from the fact that respective publication is finally in press). The image above shows a multiple pairwise alignment from BLAST of a putative inner membrane protein from Porphyromonas gingivalis. Image is small but it does not really matter – colour patches seem to be visible anyway.![Reblog this post [with Zemanta]](https://i0.wp.com/img.zemanta.com/reblog_e.png)


















Timestamped FriendFeed activity – really public “profile”
Accidentaly, I have found a simple way for obtaining a time stamp for each entry and comment any person with publicly available lifestream makes on FriendFeed (except “Likes”, which do not seem to be timestamped at all). Activity of semi-randomly choosen person during the day (summarized over couple of weeks (!)) is shown below:
FriendFeed usage during 24 hours, summarized over couple of days.
While relation between AM and PM periods is correct, time-zone is manually shifted, so it’s more difficult to guess who’s this activity is (but it’s not Robert Scoble if you want to ask). What does it tell? Basically, this person does not close FriendFeed window for the most of the day. Additionally, there’s a period of the day in which “catching-up” has place. Nothing interesting so far? Original data has much more details. It is possible for example to collect information when during the day particular person usually watches videos on YouTube. Guess – is that during working hours? 🙂
Ability to get that data for couple of weeks back without any trouble (I didn’t need to track this person’s activity for such period) was kind of disturbing. I knew it’s very simple to start tracking my habits, but I wasn’t aware of the fact that it’s also easy to see what I was doing over the last three weeks. Do you think it makes a difference?
Posted by Pawel Szczesny on January 29, 2009 in Comments, Visualization
Tags: activity tracking, Blog, FriendFeed, RSS