Modularity is one of the most interesting features of the trimeric autotransporter adhesins, and probably one of the most frustrating. As I wrote before, domain annotation is quite difficult, especially that these proteins can have often few thousands residues in length. 
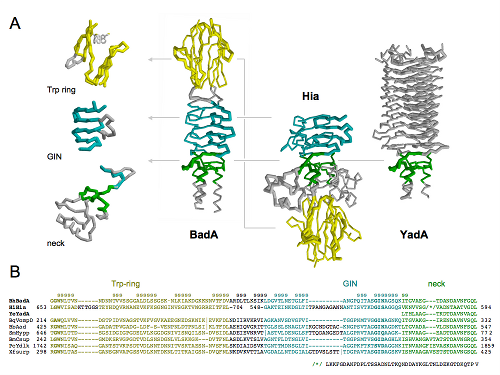
BadA, the major adhesin of Bartonella henselae, is probably the best known large TAA out there. Its sequence served us as a unofficial benchmark for domain annotation tool. Its head consist of three domains, one resembling head of YadA and two others which we claimed are similar to Hia head domains. The claim at the moment of starting this project wasn’t supported very well – Evalues of HHpred alignments were around 1 (of course all less sensitive tools didn’t see anything), but we knew they must be similar (because that two,three conserved residues were at exactly where we expected). Crystal structure of these two domains from BadA couldn’t be solved directly, so we’ve attempted molecular replacement and that worked. On the picture above you can see three known head structures for TAAs, BadA (ours), Hia and YadA (full BadA head model in on the right) and arrangement of corresponding domains in all three proteins. The whole story and lots of pretty pictures (you must see EM figures) was published today yesterday in PLoS Pathogens (OA).
Today the story isn’t so exciting as it was at the beginning. Currently HHpred easily finds domains from Hia and BadA similar with high probability – it’s an advantage of bigger database size and more mediating sequences. But I’m still pretty happy about how it went – such projects build confidence in one’s analysis skills.







Andrew Perry
August 20, 2008 at 10:41
Impressive … so if I’m understanding this correctly the similarity you originally detected using HHpred was real (ie ultimately reflected in structural similarity)… even if the latest E-values now don’t make the feat look as impressive. Nice work !
Pawel Szczesny
August 20, 2008 at 11:01
Thanks Andrew. Yes, that was the point. Looking at sequences of a single family for couple of years paid back :). There’s a certain advantage of wrapping one’s brain around an isolated problem (TAAs are pretty unique, mostly) – I could assign domains by reading the single sequence in text editor…