First major outcome of my PhD project has just appeared in the Bioinformatics (open access). It describes a system we have design to annotate specific group of bacterial proteins.
Trimeric autotransporter adhesins (TAAs) form one of the many families of bacterial surface proteins. In medically relevant species they adhere to host cells (in non-pathogenic species we don’t know what they adhere to), therefore they are considered essential virulence factors. They are autotransporters, which means that they are passing the outer membrane by themselves – C-terminal part makes a pore through which the rest of the protein goes out. In contrary to many other autotransporters, exported part is not cut but stays attached to the membrane by the C-terminal autotransport domain. TAAs are also trimeric – the pore is made of three subunits and the exported fiber is also a trimer. The last feature is pretty unique – so far it’s the only family of bacterial surface proteins which forms fibrous trimers. Interestingly, they differ in length between few hundred and five thousands residues.
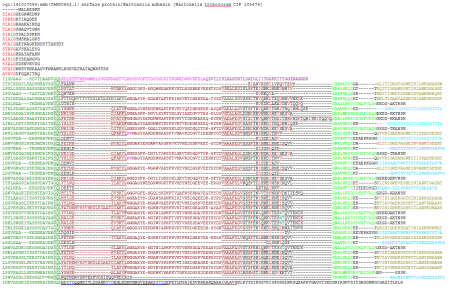
What’s so special about these proteins for bioinformatician? Structure of the fiber is not homogenous – it is a mixture of globular domains and coiled-coils. On a sequence level, they have lots of internal repeats (see the picture), heavily biased residue composition, their domain composition and architecture varies by protein. The only conserved part in all TAAs in the autotransport domain. Systems designed to identify and annotate typical protein domains (such as PFAM) don’t handle them very well – average coverage of PFAM annotation of TAAs is about 30%. The server we have built relies on the fact that domains of TAAs are exclusive for this family (they do not appear anywhere else because its unique structural constrains). Therefore we could use different thresholds, manually curated alignments and domain-context derived rules to improve the annotation.
Manual analysis of TAAs sequences is pretty tedious (well, it was, now we have the server), but on the other hand I have learnt a lot about how to read a protein sequence. I mean really read and understand how particular combination of letters influences its structure.








