One short note: I’ve started this blog with a hope that maybe I would write something useful for the next Bio::Blogs edition, which would send me first visitors. To my surprise this site was found almost within hours from the first post by Pedro Beltrao. It looks like science bloggers never sleep :).
Last year I had a chance to make a short course on protein structure prediction. One of the points I made was preparing the publication quality pictures of the models. While the Rasmol (I’m linking to open source version here) has definitely its well deserved place on the scientists computers, it is not the best choice for publication figures. My personal suggestions are listed below:
- VMD by UIUC – my favourite, steep learning curve, writes POVRay files, recent version includes Tachyon renderer and is able to use a neat feature – “ambient occlusion“
- Chimera by UCSF – pretty easy to use, recent version can render biomolecules with POVRay
- Pymol by DeLano Scientific – easy as Rasmol, has internal renderer capable producing very nice images, another favourite for completely different reasons than VMD
- Qutemol by ISTI-CNR – pretty new software and to me still in alpha state, impressive real-time rendering with ambient occlusion, capable of producing images in prof. David Goodsell style (see Molecule of the Month at PDB)
- Molscript by Avatar Software, the oldest and the most difficult to use, however the clarity of the final image is often hard to beat
Of course three first programs can do much more than just visualize the protein structure – they can be used in detailed structural analysis, can do superimpositions of protein structures, analyze trajectories from molecular simulations, display density maps, deal with alignments and many other things.
Below you can find examples of images obtained with the above software. YadA adhesin picture has an “artsy” look, but at least it shows wide range of possibilities.
 VMD
VMD
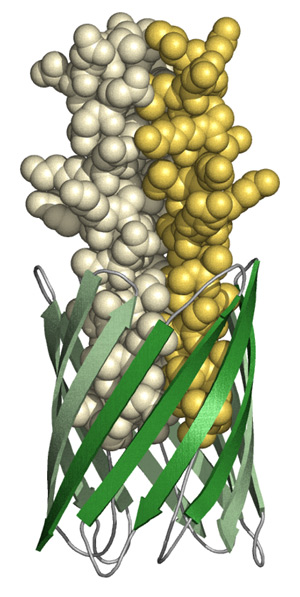 Pymol
Pymol
 Molscript
Molscript

Keith
August 7, 2007 at 21:03
Welcome Pawel, and goodluck with your new blog! Hmm.. You now have three posts including the Hello World one and you started a day after me!..Perhaps I need to add the Hello World post back in… 😉
freesci
August 7, 2007 at 21:09
It’s not the number of the post that counts 🙂 Thanks a lot.
suicyte
August 9, 2007 at 19:34
This blog has a very promising start. Good luck for the future!
I really like this comparison, some of the programs I had never heard about. Pymol and Moscript are out of the question for me, as they can only be used by academic institutions (or for big money, which I don’t have). Qutemol looks quite “qute” on the web, but it doesn’t work for me as it requires a modern graphics card (I only have the cheap on-board graphics – no luck). Chimera works, although I don’t find it that easy to use. Actually, I find it quite complicated, with very deep sub-menu structures and stuff at places where I don’t expect it to be. VMD I haven’t tried yet (but you say it has a steep learning curve while chimera is easy to use, so after my chimera experience I don’t dare to use VMD).
What I am normally using at home is an old (free) WebLab ViewerLight 3.7 version – a very nice program, very fast, easy to use but not really powerful.
freesci
August 9, 2007 at 20:13
Thanks a lot!
There are many more or less powerful programs for molecular visualization. SDSC holds large and up-to-date list of this software: http://molvis.sdsc.edu/visres/molvisfw/titles.jsp . You may want to try the BALLview. It’s one of the newest additions but it seems to be stable and quite powerful. I wasn’t very excited about it, but some of my colleagues just love this program. Have a look – maybe it just fits your needs.
VMD is a multiwindow software – it’s not very easy to use, unless one works a lot with GIMP or something similar. Also one needs to learn selection expressions syntax, while in Chimera it’s just selecting a fragment with a mouse in the sequence window… Overall I like both, and both are able to render pretty nice graphics.
Deepak
August 9, 2007 at 20:31
If I could pitch something that was released just as I was leaving Accelrys. Discovery Studio Visualizer is the next generation of WebLab ViewerLight, and significantly more powerful. Best of all, it runs on Linux. IMO, PyMol is still the best, but unfortunately not as easily accessible as it used to be.
Having used VMD through grad school (from version 1), I must admit to having a very soft spot for it. I think that it’s selection syntax is what makes it as powerful as it is.
Deepak
August 9, 2007 at 20:32
Whoops forgot to close the link. Could you please do that for me?
suicyte
August 9, 2007 at 21:24
Pawel: thanks for the information. Maybe the reason why I find Chimera difficult is that I did not find a sequence window. So far, I have done all selection by command line syntax. I will keep on looking. Other things I find non-intuitive include i) viewing options filed under “actions”, ii) simple coloring found under “actions:color”, but more sophisticated ones under “tools:structure_analysis:render_by_attribute” and surface coloring under “tools:surface_analysis:electrostatic_surface_coloring iii) there doesn’t even seem to be a built-in electrostatic surface coloring, but only a loading of external map data (which require external programs most of which cannot be used freely by people in a company. Anyway, I do like the rendered ribbon graphs and will have a look at Ballview.
Deepak: thanks for the link to DS Visualiser. Didn’t know that you used to be with Accelrys. BTW, is WLviewer an Accelrys product? It claims to be from MSI. If memory serves me right, I once tried a DSViewer, but found that I do not need the new features, and since the old WLviewer was faster on my old machine I sticked with that.
Sandra Porter
August 16, 2007 at 17:30
Hi Pawel,
What do you think about Cn3D, the viewer from the NCBI? I really like it since it has a nice GUI and is very easy for students to use, but I’m curious about what other people think.
Thanks,
Sandy
freesci
August 16, 2007 at 18:14
Dear Sandra,
I would love to try Cn3D, unfortunately their Unix version is compiled against very specific libraries and it does not work on many linux systems. The same sad story is for the Swiss PDB Viewer, which has enormous amount of bugs in its linux version (it’s almost unusable).
I don’t really understand that – software written by two major institutions handling biological data is poorly ported to other than paid operating systems, while the academic research groups or private companies have versions working on all major platforms.
Sandra Porter
August 17, 2007 at 03:53
Ah, that’s too bad. If it’s any consolation, there has been a version 4.2 in the works for about 2 years now and version 4.1 doesn’t work on VISTA either.
The person who wrote the most recent version is a friend of mine and has been really busy working on PubChem, so the release has been delayed a bit.
I can write to him and ask about versions for *nix systems but I suspect it won’t get them out any faster.
Kay at Suicyte
August 17, 2007 at 20:56
I tried Cn3D a long time ago and didn’t like it at all. I can’t really remember why, it think is was relatively fast but the structures looked kind of ugly. I guess that the program has changed since than and maybe I should try a more recent version. Anyway, what I am looking for is a program that creates decent figures for publications, but to my disappointment most programs are incapable of producing good looking output on white background. I think what is mostly lacking is the introduction of thin black lines that delimit the ribbons and other features – in the programs that can do it this really makes a hell of a difference.
With regard to spdbv: I am using this program frequently, but not for creating figures. I haven’t found another program that would offer the same (accessible) features for structural superposition and structure-based alignment generation. Sometimes I use the Linux version but I agree that it is very buggy – and I was not able to configure any reasonable fonts. Most of the problems with spdbv come probably from the history of this program. Initially, it was the private work of a single person (maybe two) who wrote the software on a Mac. I am not sure how the port to other operatinig systems has been done, but I guess that it uses some compatibility libraries. At least some of the fonts look rather Mac-like.
iayork
August 19, 2007 at 13:37
QuteMol looks really intriguing but the Mac version crashes on opening on my Macbook Pro:
Library not loaded: /sw/lib/libpng.3.dylib
Referenced from: /Users/iayork/Desktop/QuteMol.app/Contents/MacOS/qutemol
Reason: image not found
I find PyMol’s attitude (“ today’s PyMOL users can choose to either sponsor the project to enjoy incentives such as our precompiled executables, or they can choose to assume responsibility for obtaining their own FREE executables from the open-source code.“) off-putting. Of course, they can present their software any way they like, but the tone of aggrieved self-righteousness doesn’t make them look good, and it’s not like they have the only product around.
I’ve at least played with most of the other ones you list, and haven’t been blown away by any, but I’m not even at the bottom end of users. My needs (“wants”, really) are very simple, since I don’t do any real modeling; I just want to get some nice pictures, mostly for illustration purposes (in classes and so on) — so I’m interested in the “publication-quality images” part, with a simple interface on top of it.
CN3D is very easy to use and turns out OK but unexciting images. Chimera seems to have a few more options, and I think the images sometimes turn out a little nicer than CN3D, but at the simple level the difference in quality seems small.
Pedro Beltrao
August 19, 2007 at 16:26
I’ll put in another name, YASARA (www.yasara.org).
Deepak
August 19, 2007 at 17:41
YASARA looks good to. Never tried it but it was always intriguing as was VEGA which I last used about 7 years ago.
Re DSViewer … it got completely re-written a couple of years ago, so the performance should be better. It’s essentially WLViewer resurrected. It does have more features than a lot of people need though, and is probably more useful if you are doing drug design.
I have mixed feelings on PyMol. It’s the best visualization software I’ve used, but I feel Warren has lost a lot of goodwill with the current approach, especially in the academic community.
freesci
August 19, 2007 at 17:50
Iayork, in my group Molscript, Pymol and VMD have been used for publications with at least acceptable results. Advanced features of Pymol/Chimera/VMD (several layers of the visualization, like transparent surfaces and such) look very well on the screen, but unfortunately often not on the paper. I really like the output of Molscript, but it takes hours to prepare a single figure (I’ve tested various interfaces to the Molscript, but at least on the linux, I haven’t found anything that suited me). What kind of images you are trying to get?
Pedro, thanks for pointing that. I’ve used Yasara for various things (the Dynamics package), but never thought to think about it in terms of producing images. I may want to try. 🙂
freesci
August 19, 2007 at 17:57
Deepak, I’m sorry that publishing of your comment took so long – it was somehow identified by Akismet as spam :).
iayork
August 19, 2007 at 19:15
Pedro: I’ll put in another name, YASARA (www.yasara.org).
Thanks for the pointer, Pedro. I had forgotten about YASARA since I moved to my Intel Macs. Now that I can run it, I’m impressed – pretty pictures and ease of use combined.
Freesci: What kind of images you are trying to get? I’m usually after something like a PNG file (for slides or web viewing) and I’m usually trying to make pretty simple points. For example, I like to show my students MHC class I peptide complexes with the peptide’s surface-exposed surfaces and anchor residues indicated. YASARA seems to make it quite easy to make those clear.
naga
August 20, 2007 at 13:06
hi all,
pymol is good for visualization.. i liked it very much, becoz it provides the option to make molecular animation out of that..! its good wen i make presentation for my lab meets..! the ray fuction in the pymol provides a pretty good picture..!
n sector (sgi) is also used to make pretty good pictures for publications..!
naga
baoilleach
September 2, 2007 at 11:34
iayork says:
“I find PyMol’s attitude (” today’s PyMOL users can choose to either sponsor the project to enjoy incentives such as our precompiled executables, or they can choose to assume responsibility for obtaining their own FREE executables from the open-source code.“) off-putting. Of course, they can present their software any way they like, but the tone of aggrieved self-righteousness doesn’t make them look good, and it’s not like they have the only product around.”
Just to point out that PyMol is written by a single person, Warren DeLano, not a “they”. Warren is trying to maintain a delicate balance between making it open source (for example, it’s included in most major Linux distributions) and relying on it for income (you need to pay for Windows binaries, or figure out how to compile it yourself). Most of the other software discussed here (with the exception of Qutemol) is not even open source (which is why you can’t port it, fix the bugs, or build on it), unlike PyMol.
Egon Willighagen
October 29, 2007 at 20:18
Another name: Jmol.org does a pretty good PovRay export nowadays too.
Tushar
May 19, 2009 at 00:09
Can someone tell me how I will make one thick and another think loop in protein using Pymol?